 The crystal is illuminated with a finely focused monochromatic beam of X-rays, producing a diffraction pattern of regularly spaced spots known as reflections. The goniometer is used to position the crystal at selected orientations. In a single-crystal X-ray diffraction measurement, a crystal is mounted on a goniometer. X-ray crystal structures can also account for unusual electronic or elastic properties of a material, shed light on chemical interactions and processes, or serve as the basis for designing pharmaceuticals against diseases. X-ray crystallography is still the primary method for characterizing the atomic structure of new materials and in discerning materials that appear similar by other experiments. The method also revealed the structure and function of many biological molecules, including vitamins, drugs, proteins and nucleic acids such as DNA. In its first decades of use, this method determined the size of atoms, the lengths and types of chemical bonds, and the atomic-scale differences among various materials, especially minerals and alloys. Since many materials can form crystals-such as salts, metals, minerals, semiconductors, as well as various inorganic, organic, and biological molecules-X-ray crystallography has been fundamental in the development of many scientific fields. From this electron density, the mean positions of the atoms in the crystal can be determined, as well as their chemical bonds, their crystallographic disorder, and various other information. 
By measuring the angles and intensities of these diffracted beams, a crystallographer can produce a three-dimensional picture of the density of electrons within the crystal. X-ray crystallography is the experimental science determining the atomic and molecular structure of a crystal, in which the crystalline structure causes a beam of incident X-rays to diffract into many specific directions. Please discuss this issue on the article's talk page. Please consider splitting content into sub-articles, condensing it, or adding subheadings. 1953 171:737–8.This article may be too long to read and navigate comfortably. Molecular structure of nucleic acids a structure for deoxyribose nucleic acid. New CCP13 software and the strategy behind further developments: stripping and modeling of fiber diffraction data. Squire J, AL-Khayat H, Strunther A, Cranshaw J, Denny RC, Diakun G, Dover D, Forsyth VT, He A, Knupp C, Mant G, Ganeshalingam R, Rodman M, Shotton M, Windle AH. Molecular architecture in muscle contractile assemblies. Squire JM, Al-Khayat HA, Knupp C, Luther PK. Structure of the core and central channel of bacterial flagella. Plenum Publishing corporation, New York 1982 1981. Structural molecular biology: methods and applications. In: Davies DB, Srenger W, Danyluk SS, editors. CLEARER: a new tool for the analysis of X-ray fiber diffraction patterns and diffraction simulation from atomic structural models. X-ray diffraction procedures: for polycrystalline and amorphous materials. 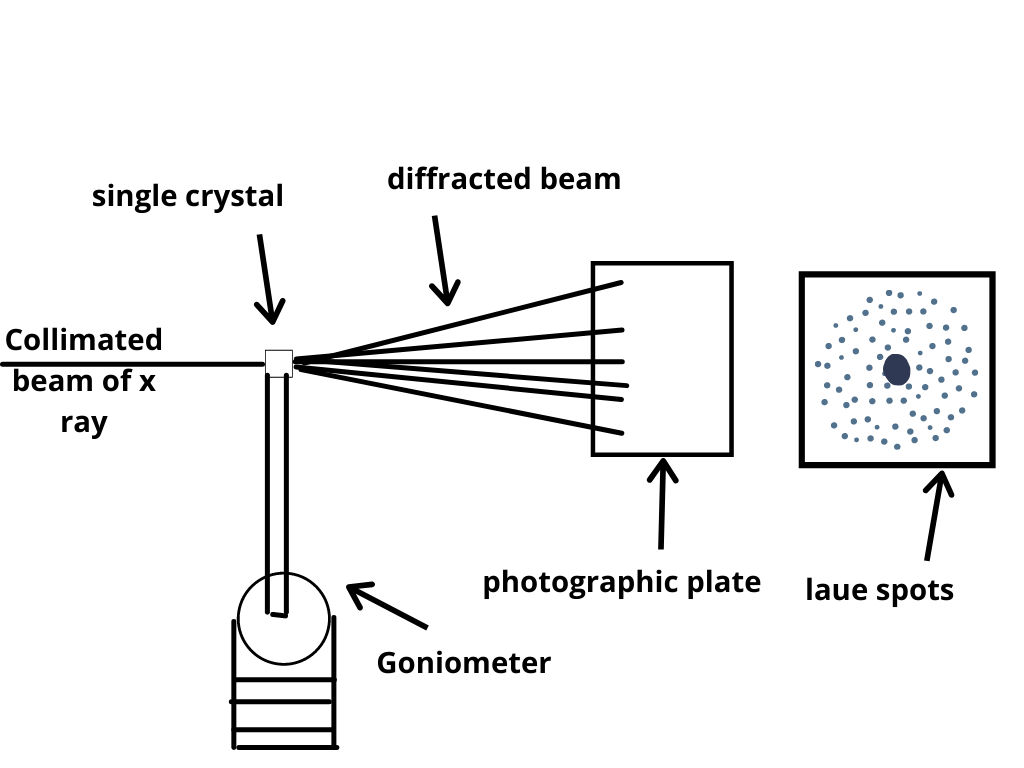
The molecular configuration of deoxyribonucleic acid.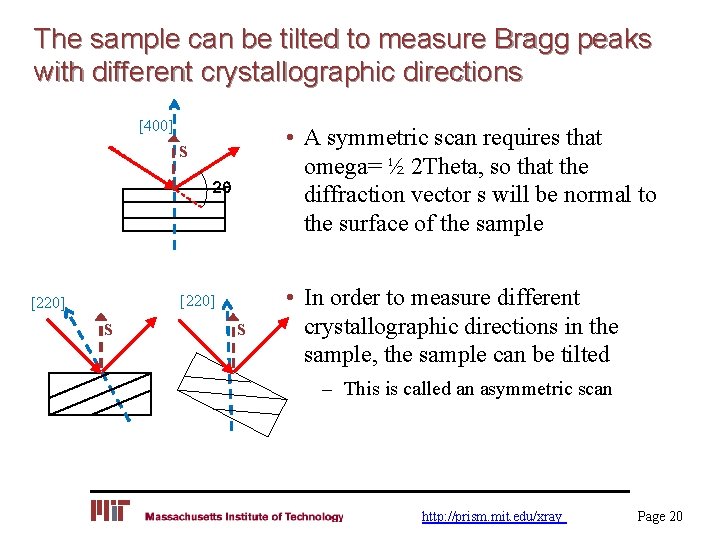 
Fuller W, Wilkins MH, Wilson HR, Hamilton LD. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
